TNNC1 in Hypertrophic Cardiomyopathy (HCM)
Evidence for role of TNNC1 in HCM
A detailed analysis of the role of non-sarcomeric genes in HCM - incorporating clinical sequencing data from the OMGL and LMM labs, a cohort of over 800 HCM probands from the Royal Brompton Hospital, London and the National Heart Centre, Singapore (sequenced on a broad cardiac NGS panel) and published sequencing, segregation and functional data - has clarified the involvement of TNNC1 variants in this condition (see our study published in the European Heart Journal for further details).
Based on this analysis, TNNC1 is classified as having: Weak Evidence
| Case excess (gene) | Max LOD score | Case excess (variant) | De novo variant | ||||
|---|---|---|---|---|---|---|---|
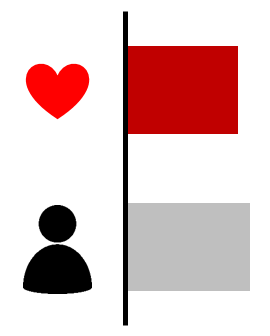 | p=0.002 | 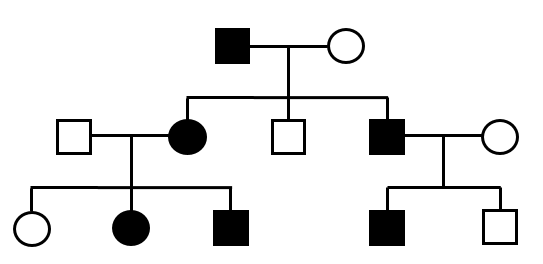 | 1.2 |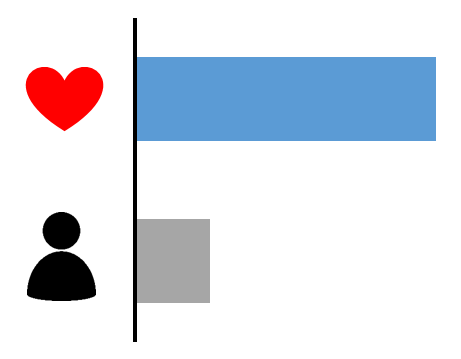  | - | 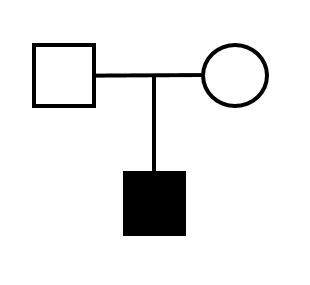 | - |
| See details below | c.363dupG (PMID:21262074) | ||||||
| Cohort (reference) | HCM patients tested | Rare variants | Case Frequency | Significance vs ExAC |
|---|---|---|---|---|
| 18572189 | 1025 | 3 | 0.00293 | p=0.027 |
| 25351510 | 874 | 3 | 0.00343 | p=0.018 |
| 27532257 | 632 | 0 | 0.00000 | no excess |
| 28082330 | 804 | 2 | 0.00249 | p=0.088 |
| Total | 3335 | 8 | 0.00240 | p=0.002 |
Summary of the frequency of rare TNNC1 variants (ExAC frequency < 0.0001) in published cohorts of HCM probands. P-values shown are from Fisher' Exact test compared to rare variants in ExAC (ExAC frequency = 0.00060). Significance is based on multiple testing correction of 31 genes tested (p<0.0016).
Data from clinical cohorts
By comparing the frequency of TNNC1 variants in large HCM clinical cohorts to the background population rate in the ExAC database, the proportion of HCM patients with pathogenic mutations in TNNC1 can be estimated, as well as the likelihood that a rare (population allele frequency <0.0001) TNNC1 variant identified in a HCM patient is disease-causing. Summary data for different variant classes (all protein-altering variants, loss of function truncating variants and non-truncating variants) is highlighted - see the table below for full details of this analysis.
Excess of TNNC1 variants in HCM: -0.06% (p=1.0000)
Based on an analysis of all rare protein-altering variants (MAF<0.0001 in ExAC) in TNNC1 found in 632 HCM samples sequenced by OMGL and LMM and in reference samples of the ExAC population database. As there is no excess burden of variation in the disease cohort, this analysis suggests that TNNC1 variants do not contribute to HCM.
Metrics by Variant Class:
| All Vars | Truncating | Non-Truncating | |
|---|---|---|---|
| Excess in HCM | -0.06% p=1.0000 |
-0.00% p=1.0000 |
-0.05% p=1.0000 |
| Etiological fraction | - | - | - |
| Odds Ratio | 0.00 0.00 - 10.27 |
0.00 0.00 - 180.09 |
0.00 0.00 - 10.89 |
The Etiological Fraction (EF) is the proportion of affected carriers where the variant caused HCM. The Odds Ratio (OR) describes the odds of having a rare variant in the patient cohort to the odds in the ExAC cohort. Fisher's exact test p-values are displayed for case excess, 95% confidence intervals for EFs and ORs.
| Source | Samples Tested | Variant Type | Variants detected | Frequency in HCM |
Frequency in ExAC | Case Excess in HCM |
|---|---|---|---|---|---|---|
| LMM2 | 632 | All | 0 | 0.00000 | 0.00060 | -0.00060 |
| Truncating | 0 | 0.00000 | 0.00004 | -0.00004 | ||
| Non-Truncating | 0 | 0.00000 | 0.00054 | -0.00054 |
References
1. Roddy Walsh, Kate L. Thomson, James S. Ware, Birgit H. Funke, Jessica Woodley, Karen J. McGuire, Francesco Mazzarotto, Edward Blair, Anneke Seller, Jenny C. Taylor, Eric V. Minikel, Exome Aggregation Consortium, Daniel G. MacArthur, Martin Farrall, Stuart A. Cook and Hugh Watkins. Reassessment of Mendelian gene pathogenicity using 7,855 cardiomyopathy cases and 60,706 reference samples. Genet Med. 2016 doi:10.1038/gim.2016.90.
2. Alfares AA, Kelly MA, McDermott G, Funke BH, Lebo MS, Baxter SB, Shen J, McLaughlin HM, Clark EH, Babb LJ, Cox SW, DePalma SR, Ho CY, Seidman JG, Seidman CE, Rehm HL. Results of clinical genetic testing of 2,912 probands with hypertrophic cardiomyopathy: expanded panels offer limited additional sensitivity. Genet Med. 2015 Nov;17(11):880-8.
3. Roddy Walsh, Rachel Buchan, Alicja Wilk, Shibu John, Leanne E. Felkin, Kate L. Thomson, Tang Hak Chiaw, Calvin Chin Woon Loong, Chee Jian Pua, Claire Raphael, Sanjay Prasad, Paul J. Barton, Birgit Funke, Hugh Watkins, James S. Ware, Stuart A. Cook. Defining the genetic architecture of hypertrophic cardiomyopathy: re-evaluating the role of non-sarcomeric genes. Eur Heart J. 2017 doi:10.1093/eurheartj/ehw603.